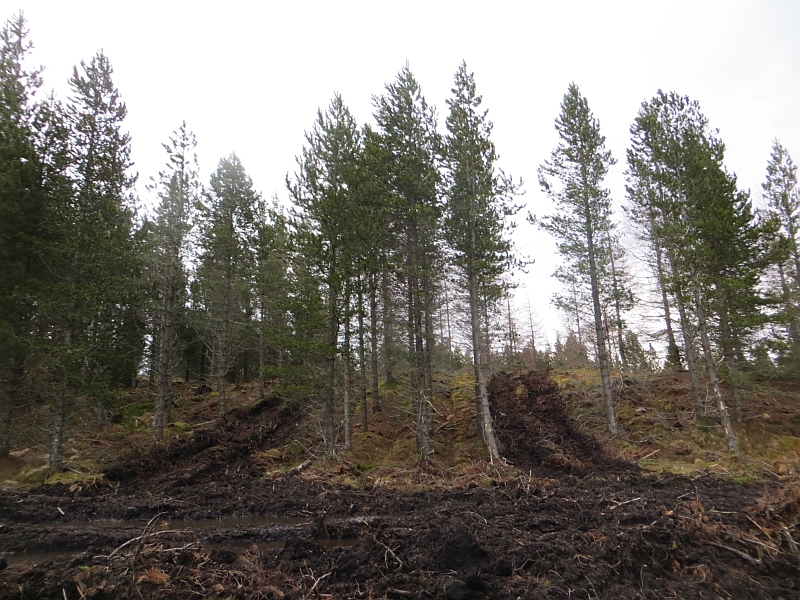
The major threats to the sustainable supply of forest tree products are adverse climate, pests, and diseases. Climate change, exemplified by increased drought, poses a unique threat to global forest health. This is attributed to the unpredictable behavior of forest pathosystems, which can favor fungal pathogens over the host under persistent drought stress conditions in the future. Currently, the effects of drought on tree resistance against pathogens are hypothetical, thus research is needed to identify these correlations. Norway spruce (Picea abies (L.) H. Karst.) is one of the most economically important tree species in Europe and is considered highly vulnerable to changes in climate. Dedicated experiments to investigate how disturbances will affect the Norway spruce—Heterobasidion sp. pathosystem is important, in order to develop different strategies to limit the spread of H. annosum s.l. under the predicted climate change. Here, we report a transcriptional study to compare Norway spruce gene expressions to evaluate the effects of water availability and the infection of Heterobasidion parviporum. We performed inoculation studies of three-year-old saplings in a greenhouse (purchased from a nursery). Norway spruce saplings were treated in either high (+) or low (−) water groups: the high water group received double the water amount than the low water group. RNA was extracted and sequenced. Similarly, we quantified gene expression levels of candidate genes in biotic stress and jasmonic acid (JA) signaling pathways using qRT-PCR, through which we discovered a unique preferential defense response of H. parviporum-infected Norway spruce under drought stress at the molecular level. Disturbances related to water availability, especially low water conditions can have negative effects on the tree host and benefit the infection ability of the pathogens in the host. From our RNA-seq analysis, 114 differentially expressed gene regions were identified between high (+) and low (−) water groups under pathogen attack. None of these gene pathways were identified to be differentially expressed from both non-treated and mock-control treatments between high (+) and low (−) water groups. Finally, only four genes were found to be associated with drought in all treatments.
Drought stress described by transcriptional responses of Picea Abies (L.) H. Karst. under Pathogen Heterobasidion parviporum Attack
Year: 2021